Thanks, Mufei.
If you have time, it is appreiated that you could take a look of my implementation of Message Passing with Edges Update. I know it is quite impolite to request you to debug for me. Thus if you do not have time, please forget this.
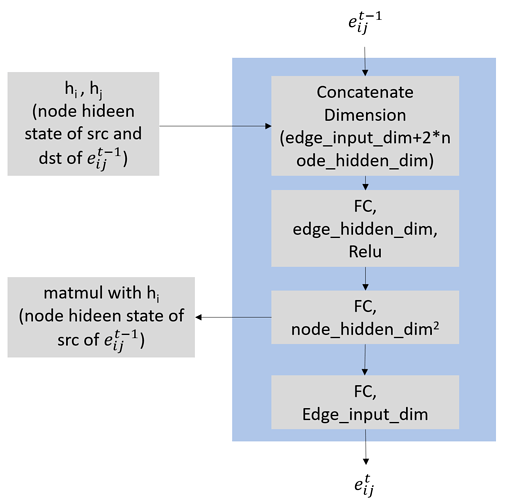
The edge update function I want to use is something like below
However, when I added this into the MPNN model of
@geekinglcq , it does not work.
#!/usr/bin/env python
# coding: utf-8
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.nn import Parameter
from torch.utils.data import dataloader
import dgl
import dgl.function as fn
import dgl.nn.pytorch as dgl_nn
import os
from os import listdir
from os.path import isfile, join
import networkx as nx
import numpy as np
import pickle
class NNConvLayer(nn.Module):
"""MPEU Conv Layer
Paramters
---------
"""
def __init__(self,
in_channels,
out_channels,
edge_net,
edge_lin,
root_weight=True,
bias=True):
super(NNConvLayer, self).__init__()
self.in_channels = in_channels
self.out_channels = out_channels
self.edge_net = edge_net
self.edge_lin =edge_lin
if root_weight:
self.root = Parameter(torch.Tensor(in_channels, out_channels))
else:
self.register_parameter('root', None)
if bias:
self.bias = Parameter(torch.Tensor(out_channels))
else:
self.register_parameter('bias', None)
self.reset_parameters()
def reset_parameters(self):
if self.root is not None:
nn.init.xavier_normal_(self.root.data, gain=1.414)
if self.bias is not None:
self.bias.data.zero_()
for m in self.edge_net.modules():
if isinstance(m, nn.Linear):
nn.init.xavier_normal_(m.weight.data, gain=1.414)
def message(self, edges):
return {'m': torch.matmul(edges.src['h'].unsqueeze(1), edges.data['w']).squeeze(1)}
def apply_node_func(self, nodes):
aggr_out = nodes.data['aggr_out']
if self.root is not None:
aggr_out = torch.mm(nodes.data['h'], self.root) + aggr_out
if self.bias is not None:
aggr_out = aggr_out + self.bias
return {'h': aggr_out}
def update_edge(self, edges):
cedge = torch.cat((edges.src['h'], edges.dst['h'], edges.data['e_feat']), -1)
#cedge = cedge.unsqueeze(-1) if cedge.dim() == 1 else cedge
cedge = self.edge_net(cedge)
wedge = cedge.view(-1, self.in_channels, self.out_channels)
nedge = self.edge_lin(cedge)
return {"w": wedge, "e_feat":nedge}
def forward(self, g, h, e):
h = h.unsqueeze(-1) if h.dim() == 1 else h
e = e.unsqueeze(-1) if e.dim() == 1 else e
g.ndata['h'] = h
g.apply_edges(self.update_edge)
g.update_all(self.message, fn.sum("m", "aggr_out"), self.apply_node_func)
return g.ndata.pop('h')
class MPEUModel(nn.Module):
"""MPEU model"""
def __init__(self,
node_input_dim=15,
edge_input_dim=5,
output_dim=12,
node_hidden_dim=64,
edge_hidden_dim=128,
num_step_message_passing=6,
num_step_set2set=6,
num_layer_set2set=3,
device=torch.device("cuda")):
"""model parameters setting
Paramters
---------
"""
super(MPEUModel, self).__init__()
self.name = "MPEU"
self.num_step_message_passing = num_step_message_passing
self.lin0 = nn.Linear(node_input_dim, node_hidden_dim)
self.device = device
edge_network = nn.Sequential(
nn.Linear(int(edge_input_dim+2*node_hidden_dim), edge_hidden_dim),
nn.ReLU(),
nn.Linear(edge_hidden_dim, node_hidden_dim * node_hidden_dim)
)
edge_lin = nn.Linear(node_hidden_dim * node_hidden_dim, edge_input_dim)
self.conv = NNConvLayer(in_channels=node_hidden_dim, out_channels=node_hidden_dim, edge_net=edge_network,edge_lin = edge_lin,
root_weight=False)
self.gru = nn.GRU(node_hidden_dim, node_hidden_dim)
self.set2set = dgl_nn.glob.Set2Set(node_hidden_dim, num_step_set2set, num_layer_set2set)
self.lin1 = nn.Linear(2 * node_hidden_dim, node_hidden_dim)
self.lin2 = nn.Linear(node_hidden_dim, output_dim)
def forward(self, g):
h = g.ndata['n_feat']
#print(h)
h = h.to(self.device)
out = F.relu(self.lin0(h))
h = out.unsqueeze(0)
for i in range(self.num_step_message_passing):
h = self.conv(g, out, g.edata['e_feat'])
m = F.relu(h)
out, h = self.gru(m.unsqueeze(0), h.unsqueeze(0))
out = out.squeeze(0)
out = self.set2set(g, out)
out = F.relu(self.lin1(out))
out = self.lin2(out)
return out
I felt like this should be some problem of the dgl graph size during forwarding, but I have no idea which one currently. Thanks in advance.