Hi all,
Question: What is the proper way of storing embeddings outside of GPU and only loading to GPU the ones beings used, without repetition, during training?
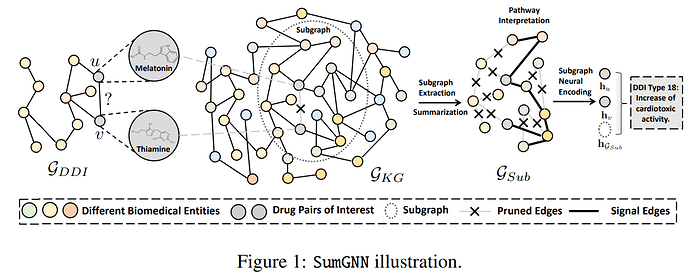
Context: I am using a biological knowledge graph. When I am predicting something like a GENE-DRUG interaction, I sample a n-hops subgraph around this node pair, and then classify the subgraph. My goal is to do graph batching. I am starting with 1-hop subgraph, which usually has around 600 nodes and 30k edges. The whole graph has over 300.000 nodes.
Lets suppose that my batch has 2 subgraphs of 600 nodes each, and they have 100 nodes in common. Where do I store the embeddings so that backpropagation updates the embeddings properly and GPU memory is not wasted?
- In the node?
- In the dataloader?
- In the model?
Thanks you so much for your time! I have been reading the documentation for a while and cant figure it out. If it is not a big deal, would you please write a quick example like the one below?
import dgl.nn.pytorch as dglnn
import torch.nn as nn
import torch
class Classifier(nn.Module):
def __init__(self, in_dim, hidden_dim, n_classes):
super(Classifier, self).__init__()
self.conv1 = dglnn.GraphConv(in_dim, hidden_dim)
self.conv2 = dglnn.GraphConv(hidden_dim, hidden_dim)
self.classify = nn.Linear(hidden_dim, n_classes)
def forward(self, g, h):
# Apply graph convolution and activation.
h = F.relu(self.conv1(g, h))
h = F.relu(self.conv2(g, h))
with g.local_scope():
g.ndata['h'] = h
# Calculate graph representation by average readout.
hg = dgl.mean_nodes(g, 'h')
return self.classify(hg)
import dgl.data
dataset = dgl.data.GINDataset('MUTAG', False)
from dgl.dataloading import GraphDataLoader
dataloader = GraphDataLoader(
dataset,
batch_size=1024,
drop_last=False,
shuffle=True)
import torch.nn.functional as F
# Only an example, 7 is the input feature size
model = Classifier(7, 20, 5)
opt = torch.optim.Adam(model.parameters())
for epoch in range(20):
for batched_graph, labels in dataloader:
feats = batched_graph.ndata['attr']
logits = model(batched_graph, feats)
loss = F.cross_entropy(logits, labels)
opt.zero_grad()
loss.backward()
opt.step()